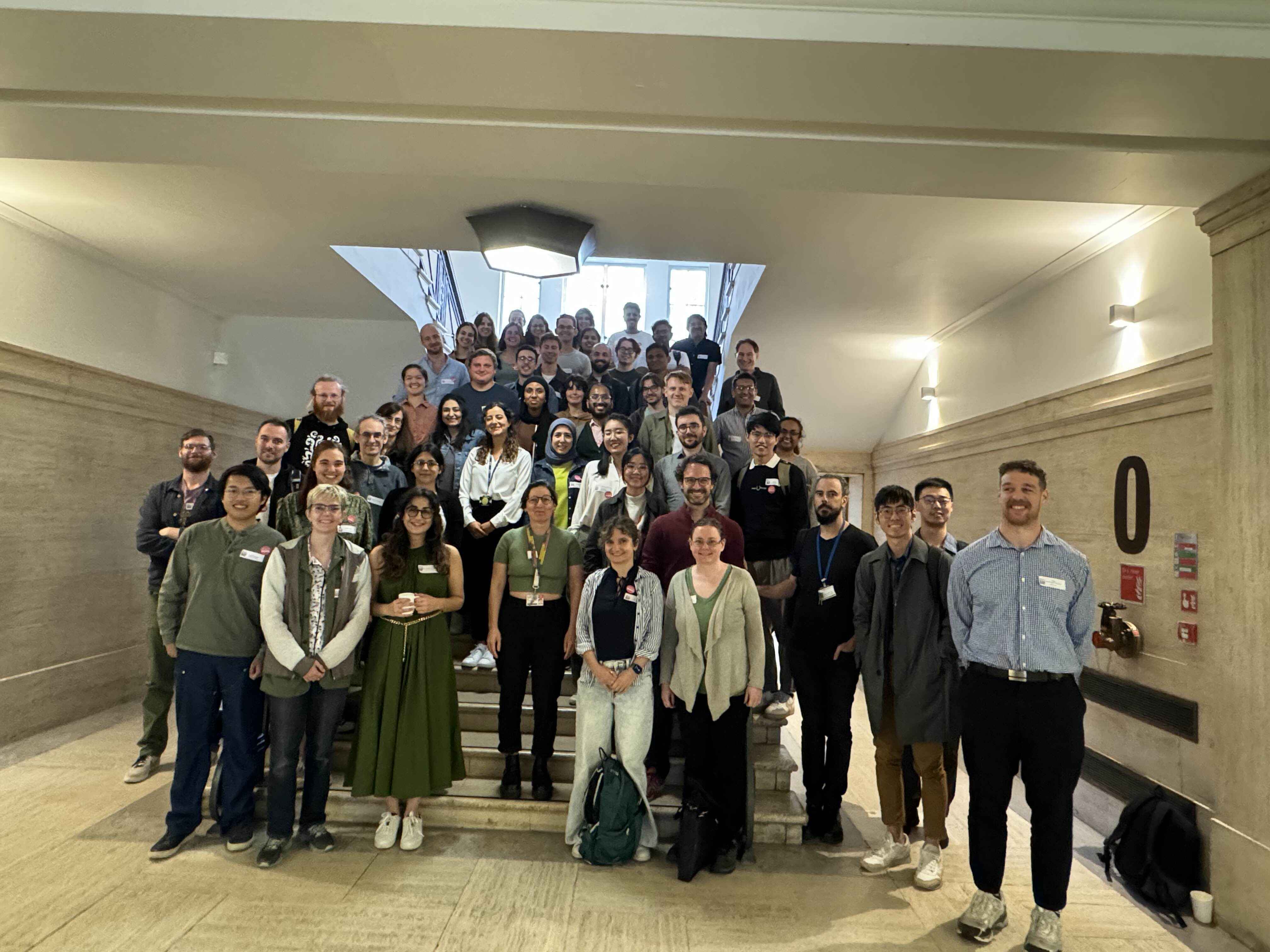
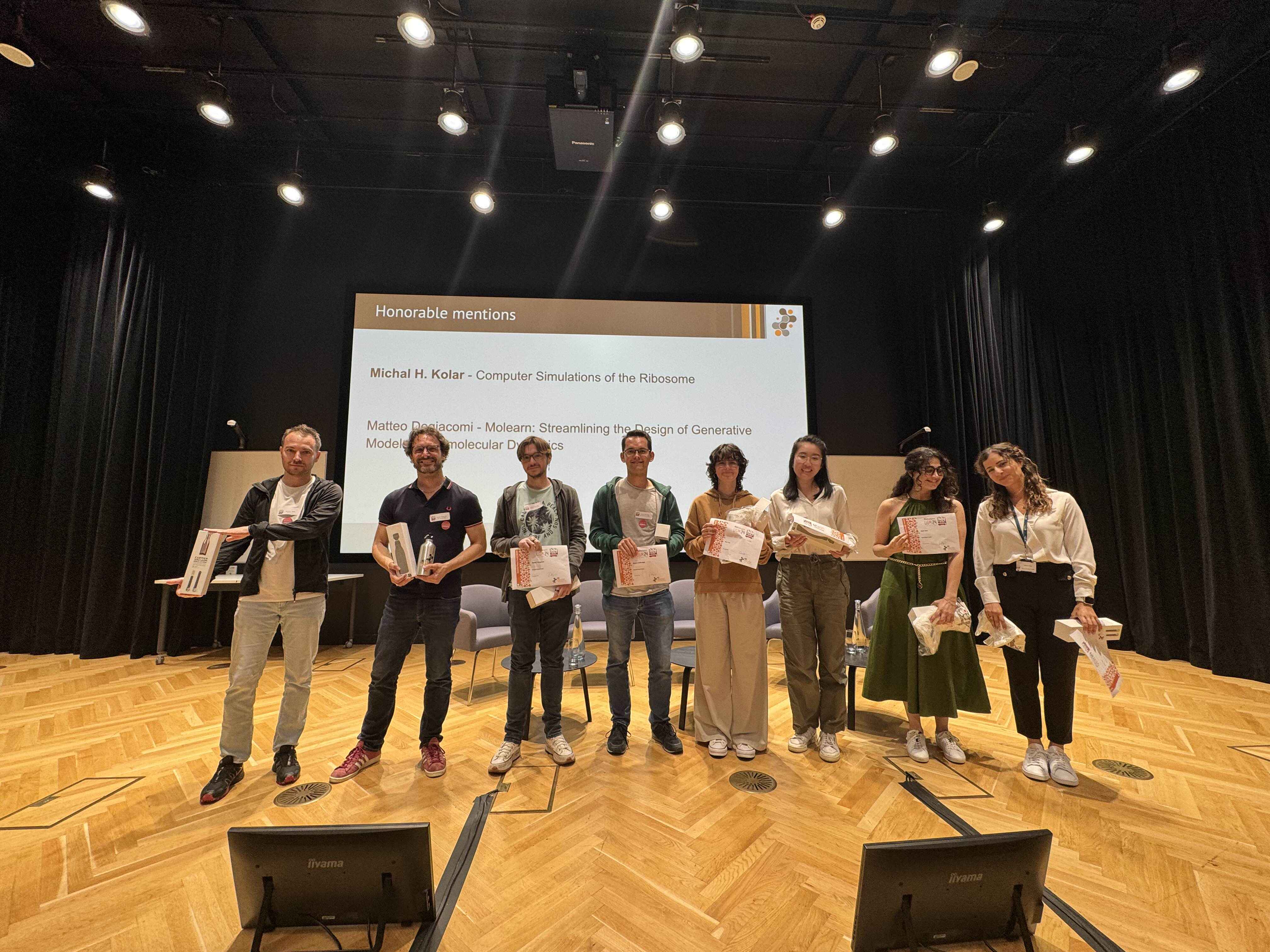
Release 2.8.0 of MDAnalysis
22 Nov 2024We are happy to release version 2.8.0 of MDAnalysis!
This is a minor release of the MDAnalysis library, which means that it contains enhancements, bug fixes, deprecations, and other backwards-compatible changes.
However, in this case minor does not quite do justice to what is happening in this release, given that we have (at least) three big changes/additions:
- The license was changed to the GNU Lesser General Public License so that MDAnalysis can be used by packages under any license while keeping the source code itself free and protected.
-
We introduce the Guesser API for guessing missing topology attributes such as element or mass in a context-dependent manner. Until release 3.0, you should not notice any differences but under the hood we are getting ready to make it easier to work with simulations in a different context (e.g., with the MARTINI force field or experimental PDB files). With consistent attributes, such as elements, it becomes a lot easier to interface with tools like the cheminformatics RDKit (via the converters).
The guessers are the GSoC 2022 project of @aya9aladdin with help from @lilyminium, @IAlibay, and @jbarnoud.1
-
We are introducing parallel analysis for tools in MDAnalysis.analysis following the simple split-apply-combine paradigm that we originally prototyped in PMDA 2. What’s really exciting is that any analysis code that is based on MDAnalysis.analysis.base.AnalysisBase can enable parallelization with a few lines of extra code—all the hard work is done behind the scenes in the base class (in a way that is fully backwards compatible!).
This new feature is the work of @marinegor who brought his GSoC 2023 project to completion, with great contributions by @p-j-smith, @yuxuanzhuang and @RMeli 3.
Not all MDAnalysis analysis classes have parallelization enabled yet but @talagayev has been working tirelessly on already updating
GNMAnalysis,BAT,Dihedral,Ramachandran,Janin,DSSP(yes, MDAnalysis has finally got DSSP, based on pydssp, also thanks to @marinegor),HydrogenBondAnalysis, in addition toRMSD.
Read on for more details on the license change and the usual information on supported environments, upgrading your version of MDAnalysis, and a summary of the most important changes.
License change to LGPL
This is the first release of MDAnalysis under the Lesser General Public License. We have been working towards this license change for the last 3 years; this release (almost) concludes the process that we described in our licensing update blog post.
- All code is now under LGPLv2.1 license or any higher version.
- The package is under the LGPLv3 license or any higher version. However, once we have removed dependencies that prevent licensing under LGPLv2.1+ at the moment, we will also license the package under the same LGPLv2.1+ as the code itself.
We would like to thank all our contributors who granted us permission to change the license. We would also like to thank a number of institutions who were especially supportive of our open source efforts, namely Arizona State University, Australian National University, Johns Hopkins University, and the Open Molecular Science Foundation. We are also grateful to NumFOCUS for legal support. The relicensing team was lead by @IAlibay and @orbeckst.
Supported environments
The minimum required NumPy version is 1.23.3; MDAnalysis now builds against NumPy 2.0.
Supported Python versions: 3.10, 3.11, 3.12, 3.13. Support for version 3.13 has been added in this release and support for 3.9 has been dropped (following SPEC 0).
Please note that Python 3.13 is limited to PyPi for now, the conda-forge channel installs only provide support for Python 3.10 to 3.12.
Supported Operating Systems:
Upgrading to MDAnalysis version 2.8.0
To update with mamba (or conda) from the conda-forge channel run
mamba update -c conda-forge mdanalysis
To update from PyPi with pip run
python -m pip install --upgrade MDAnalysis
For more help with installation see the installation instructions in the User Guide. Make sure you are using a Python version compatible with MDAnalysis before upgrading (Python >= 3.10).
Notable changes
For a full list of changes, bugfixes and deprecations see the CHANGELOG.
Enhancements:
- Added
guess_TopologyAttrs()API to the Universe to handle attribute guessing (PR #3753) - Added the
DefaultGuesserclass, which is a general-purpose guesser with the same functionalities as the existing guesser.py methods (PR #3753) - Introduce parallelization API to
AnalysisBaseand toanalysis.rms.RMSDclass (Issue #4158, PR #4304) - Add
analysis.DSSPmodule for protein secondary structure assignment, based on pydssp - Improved performance of PDBWriter (Issue #2785, PR #4472)
- Added parsing of arbitrary columns of the LAMMPS dump parser. (Issue #3504)
- Implement average structures with iterative algorithm from DOI 10.1021/acs.jpcb.7b11988. (Issue #2039, PR #4524)
- Add support for TPR files produced by Gromacs 2024.1 (PR #4523)
Fixes:
- Fix Bohrium (Bh) atomic mass in tables.py (PR #3753)
- Catch higher dimensional indexing in GroupBase & ComponentBase (Issue #4647)
- Do not raise an Error reading H5MD files with datasets like
observables/<particle>/<property>(part of Issue #4598, PR #4615) - Fix failure in double-serialization of TextIOPicklable file reader. (Issue #3723, PR #3722)
- Fix failure to preserve modification of coordinates after serialization, e.g. with transformations (Issue #4633, PR #3722)
- Fix PSFParser error when encountering string-like resids (Issue #2053, Issue #4189 PR #4582)
- Convert openmm Quantity to raw value for KE and PE in OpenMMSimulationReader.
- Atomname methods can handle empty groups (Issue #2879, PR #4529)
- Fix bug in PCA preventing use of
frames=...syntax (PR #4423) - Fix
analysis/diffusionmap.pyiteration through trajectory to iteration overself._sliced_trajectory, hence supportingDistanceMatrix.run(frames=...)(PR #4433)
Changes:
- Relicense code contributions from GPLv2+ to LGPLv2.1+ and the package from GPLv3+ to LGPLv3+ (PR #4794)
- only use distopia < 0.3.0 due to API changes (Issue #4739)
- The
fetch_mmtfmethod has been removed as the REST API service for MMTF files has ceased to exist (Issue #4634) - MDAnalysis now builds against numpy 2.0 rather than the minimum supported numpy version (PR #4620)
Deprecations:
- Deprecations of old guessing functionality (in favor of the new
Guesser API)
-
MDAnalysis.topology.guessersis deprecated in favour of the new Guessers API and will be removed in version 3.0 (PR #4752) - The
guess_bonds,vdwradii,fudge_factor, andlower_boundkwargs are deprecated for bond guessing during Universe creation. Instead, pass("bonds", "angles", "dihedrals")intoto_guessorforce_guessduring Universe creation, and the associatedvdwradii,fudge_factor, andlower_boundkwargs intoGuessercreation. Alternatively, ifvdwradii,fudge_factor, andlower_boundare passed intoUniverse.guess_TopologyAttrs, they will override the previous values of those kwargs. (Issue #4756, PR #4757) -
MDAnalysis.topology.tablesis deprecated in favour ofMDAnalysis.guesser.tablesand will be removed in version 3.0 (PR #4752) - Element guessing in the
ITPParseris deprecated and will be removed in version 3.0 (Issue #4698) -
Unknown masses are still set to 0.0 for current version, this
will be changed in version 3.0.0 and replaced by
Masses“no_value_label” attribute (np.nan) (PR #3753)
-
-
A number of analysis modules have been moved into their own MDAKits, following the 3.0 roadmap towards a trimmed down core library. Until release 3.0, these modules are still available through
MDAnalysis.analysis(either as an import of the MDAKit as an automatically installed dependency of the MDAnalysis package or as the original code) but from 3.0 onwards, users must install the MDAKit explicitly and then import it by themselves.- The
MDAnalysis.analysis.encoremodule has been deprecated in favour of the mdaencore MDAKit and will be removed in version 3.0.0 (PR #4737) - The
MDAnalysis.analysis.waterdynamicsmodule has been deprecated in favour of the waterdynamics MDAKit and will be removed in version 3.0.0 (PR #4404) - The
MDAnalysis.analysis.psamodule has been deprecated in favour of the PathSimAnalysis MDAKit and will be removed in version 3.0.0 (PR #4403)
- The
- The MMTF Reader is deprecated and will be removed in version 3.0 as the MMTF format is no longer supported (Issue #4634).
Author statistics
This release was the work of 22 contributors, 10 of which are new contributors.
Our new contributors are:
- @aditya292002
- @pstaerk
- @kainszs
- @SampurnaM
- @leonwehrhan
- @PythonFZ
- @kurtmckee
- @laksh-krishna-sharma
- @MattTDavies,
- @talagayev
Acknowledgements
MDAnalysis thanks NumFOCUS for its continued support as our fiscal sponsor and the Chan Zuckerberg Initiative for supporting MDAnalysis under EOSS4 and EOSS5 awards.
— @IAlibay (release manager) on behalf of the MDAnalysis Team
-
The Guesser API was a big undertaking: Her merged PR #3753 totalled 668 (!) comments. We look forward to the community providing Guessers for specific context. ↩
-
Shujie Fan, Max Linke, Ioannis Paraskevakos, Richard J. Gowers, Michael Gecht, and Oliver Beckstein. PMDA - Parallel Molecular Dynamics Analysis. In Chris Calloway, David Lippa, Dillon Niederhut, and David Shupe, editors, Proceedings of the 18th Python in Science Conference, 134 – 142. Austin, TX, 2019. SciPy. doi:10.25080/Majora-7ddc1dd1-013. ↩
-
Adding parallelization in a transparent manner was quite a difficult undertaking that touched many parts of the analysis code and required a lot of thought. Feedback is very welcome! See PR #4162 with 713 (!) comments. ↩